Current state: Accepted
ISSUE: https://github.com/milvus-io/milvus/issues/15604
PRs:
Keywords: bulk load, import
Released: with Milvus 2.1
Authors: yhmo
Summary
Import data by a shortcut to get better performance compared with insert().
Motivation
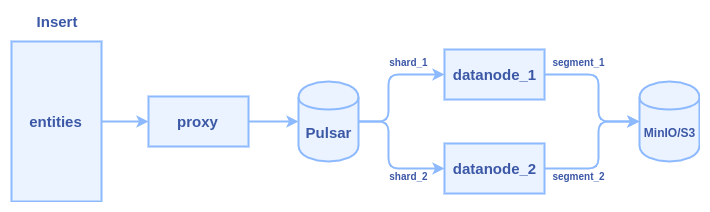
- client calls insert() to transfer data to Milvus proxy node
- proxy split the data in do multiple parts according to sharding number of the collection
- proxy constructs a message packet for each part and send the message packets into the Pulsar service
- data nodes pull the message packets from the Pulsar service, each data node pull a packet
- data nodes persist data into segment when flush() action is triggered
Typically, it cost several hours to insert one billion entities with 128-dimensional vectors. Lots of time is wasted in two major areas: network transmission and Pulsar management.
We can see there are at least three times network transmission in the process: 1) client => proxy 2) proxy => pulsar 3) pulsar => data node
We need a new interface to do bulk load without network bandwidth wasting and skip the Pulsar management. Brief requirements of the new interface:
- import data from JSON format files. (first stage)
- import data from Numpy format files. (first stage)
- copy a collection within one Milvus 2.0 service. (second stage)
- copy a collection from one Milvus 2.0 service to another. (second stage)
- import data from Milvus 1.x to Milvus 2.0 (third stage)
- parquet/faiss/csv files (TBD)
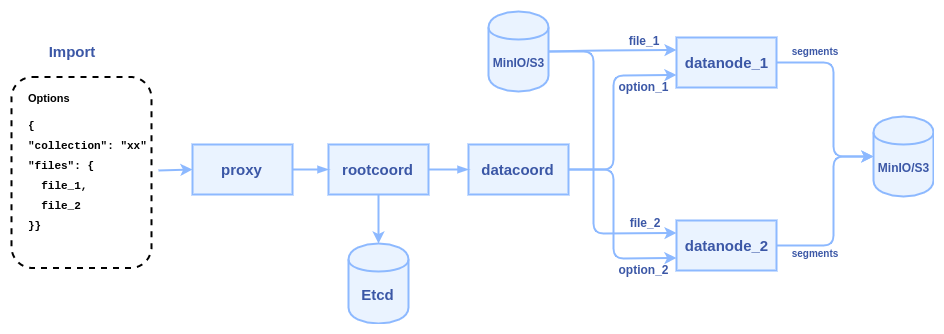
To reduce network transmission and skip Plusar management, the new interface will allow users to input the path of some data files(json, numpy, etc.) on MinIO/S3 storage, and let the data nodes directly read these files and parse them into segments. The internal logic of the process becomes:
1. client calls import() to pass some file paths to Milvus proxy node
2. proxy node passes the file paths to root coordinator, then root coordinator passes to data coordinator
3. data coordinator node picks a data node or multiple data nodes (according to the sharding number) to parse files, each file can be parsed into a segment or multiple segments.
4. once a task is finished, data node report to data coordinator, and data coordinator report to root coordinator, the generated segments will be sent to index node to build index
5. the root coordinator will record a task list in Etcd, after the generated segments successfully build index, root coordinator marks the task as "completed"
1. SDK Interfaces
The python API declaration:
def import(collection_name, files, row_based, partition_name=None, options=None)
- collection_name: the target collection name (required)
- partition_name: target partition name (optional)
- row_based: a boolean to specify row-based or column-based
- files: a list of files with row-based format or column-based format files (required)
row-based files: ["file_1.json", "file_2.json"]
column-based files: ["file_1.json", "vectors.npy"] - options: extra options in JSON format, for example: the MinIO/S3 bucket where the files come from (optional)
{"bucket": "mybucket"} - return a list of task ids
def get_import_state(task_id)
- task_id: id of an import task returned by import()
- return {state: string, row_count: integer, progress: float, failed_reason: string, id_list: list, file: string}
Note: the "state" could be "pending", "started", "downloaded", "parsed", "persisted", "completed", "failed"
Pre-defined format for import files
Assume we have a collection with 2 fields(one primary key and one vector field) and 5 rows:
| uid | vector |
|---|---|
| 101 | [1.1, 1.2, 1.3, 1.4] |
| 102 | [2.1, 2.2, 2.3, 2.4] |
| 103 | [3.1, 3.2, 3.3, 3.4] |
| 104 | [4.1, 4.2, 4.3, 4.4] |
| 105 | [5.1, 5.2, 5.3, 5.4] |
There are two ways to represent the collection with data files:
(1) Row-based data file, a JSON file contains multiple rows.
file_1.json:
{
"rows":[
{"uid": 101, "vector": [1.1, 1.2, 1.3, 1.4]},
{"uid": 102, "vector": [2.1, 2.2, 2.3, 2.4]},
{"uid": 103, "vector": [3.1, 3.2, 3.3, 3.4]},
{"uid": 104, "vector": [4.1, 4.2, 4.3, 4.4]},
{"uid": 105, "vector": [5.1, 5.2, 5.3, 5.4]}
]
}
Call import() to import the file:
import(collection_name="test", row_based = true, files=["file_1.json"])
(2) Column-based data file, a JSON file contains multiple columns.
file_1.json for the "uid" field and "vector" field:
{
"uid": [101, 102, 103, 104, 105]
"vector": [[1.1, 1.2, 1.3, 1.4], [2.1, 2.2, 2.3, 2.4], [3.1, 3.2, 3.3, 3.4], [4.1, 4.2, 4.3, 4.4], [5.1, 5.2, 5.3, 5.4]]
}
Call import() to import the file:
import(collection_name="test", row_based=false, files=["file_1.json"])
We also support store vectors in a Numpy file, we require the numpy file's name is equal to the filed name. Let's say the "vector" field is stored in vector.npy, the "uid" field is stored in "file_1.json", then we can call import():
import(collection_name="test", row_based=false, files=["file_1.json", "vector.npy"])
Note: for column-based, we don't support multiple json files, all columns should be stored in one json file. If user use a numpy file to store vector field, then other scalar fileds should be stored in one json file.
Error handling
The Import():
- Return error "Collection doesn't exist" if the target collection doesn't exist
- Return error "Partition doesn't exist" if the target partition doesn't exist
- Return error "Bucket doesn't exist" if the target bucket doesn't exist
- Return error "File list is empty" if the files list is empty
- ImportTask pending list has limit size, if a new import request exceed the limit size, return error "Import task queue max size is xxx, currently there are xx pending tasks. Not able to execute this request with x tasks."
The get_import_state():
- Return error "File xxx doesn't exist" if could not open the file.
- All fields must be presented, otherwise, return the error "The field xxx is not provided"
- For row-based json files, return "not a valid row-based json format, the key rows not found" if could not find the "rows" node
- For column-based files, if a vector field is duplicated in numpy file and json file, return the error "The field xxx is duplicated
- Return error "json parse error: xxxxx" if encounter illegal json format
- The row count of each field must be equal, otherwise, return the error "Inconsistent row count between field xxx and xxx". (all segments generated by this file will be abandoned)
- If a vector dimension doesn't equal to field schema, return the error "Incorrect vector dimension for field xxx". (all segments generated by this file will be abandoned)
- If a data file size exceed 1GB, return error "Data file size must be less than 1GB"
- If an import task is no response for more than 6 hours, it will be marked as failed
- If datanode is crashed or restarted, the import task on it will be marked as failed
2. Proxy RPC Interfaces
service MilvusService {
rpc Import(ImportRequest) returns (ImportResponse) {}
rpc GetImportState(GetImportStateRequest) returns (GetImportStateResponse) {}
}
message ImportRequest {
string collection_name = 1; // target collection
string partition_name = 2; // target partition
bool row_based = 3; // the file is row-based or column-based
repeated string files = 4; // file paths to be imported
repeated common.KeyValuePair options = 5; // import options, bucket, etc.
}
message ImportResponse {
common.Status status = 1;
repeated int64 tasks = 2; // id array of import tasks
}
message GetImportStateRequest {
int64 task = 1; // id of an import task
}
enum ImportState {
ImportPending = 0;
ImportFailed = 1;
ImportDownloaded = 2;
ImportParsed = 3;
ImportPersisted = 4;
ImportCompleted = 5;
}
message GetImportStateResponse {
common.Status status = 1;
ImportState state = 2; // is this import task finished or not
int64 row_count = 3; // if the task is finished, this value is how many rows are imported. if the task is not finished, this value is how many rows are parsed. return 0 if failed.
repeated int64 id_list = 4; // auto generated ids if the primary key is autoid
repeated common.KeyValuePair infos = 5; // more informations about the task, progress percent, file path, failed reason, etc.
}
3. Rootcoord RPC interfaces
The declaration of import API in rootcoord RPC:
service RootCoord {
rpc Import(milvus.ImportRequest) returns (milvus.ImportResponse) {}
rpc GetImportState(milvus.GetImportStateRequest) returns (milvus.GetImportStateResponse) {}
rpc ReportImport(ImportResult) returns (common.Status) {}
}
message ImportResult {
common.Status status = 1;
int64 task_id = 2; // id of the task
common.ImportState state = 3; // state of the task
repeated int64 segments = 4; // id array of new sealed segments
repeated int64 auto_ids = 5; // auto-generated ids for auto-id primary key
int64 row_count = 6; // how many rows are imported by this task
repeated common.KeyValuePair infos = 7; // more informations about the task, file path, failed reason, etc.
}
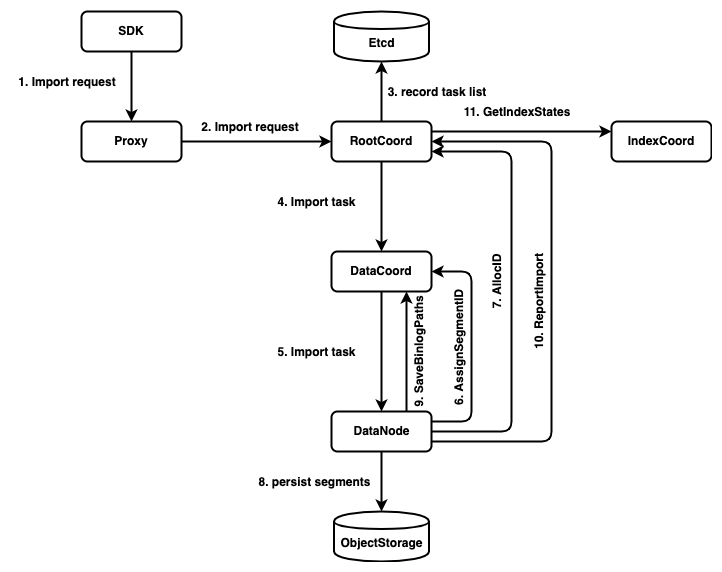
The call chain of import worflow:
4. Datacoord RPC interfaces
The declaration of import API in datacoord RPC:
service DataCoord {
rpc Import(ImportTask) returns (ImportTaskResponse) {}
}
message ImportTask {
common.Status status = 1;
string collection_name = 2; // target collection
string partition_name = 3; // target partition
bool row_based = 4; // the file is row-based or column-based
int64 task_id = 5; // id of the task
repeated string files = 6; // file paths to be imported
repeated common.KeyValuePair infos = 7; // more informations about the task, bucket, etc.
}
message ImportTaskResponse {
common.Status status = 1;
int64 datanode_id = 2; // which datanode takes this task
}
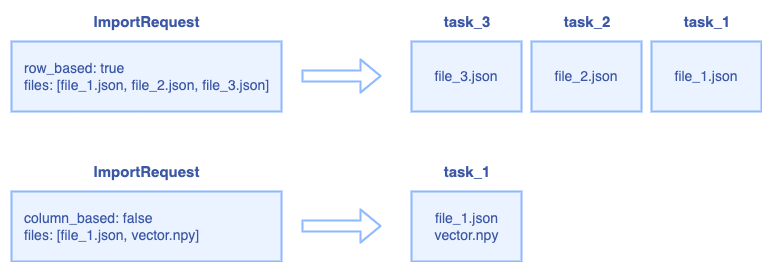
The relationship between ImportRequest and ImportTask:
For row-based request, the RootCoord splits the request into multiple ImportTask, each json file is a ImportTask.
For column-based request, all files will be regarded as one ImportTask.
5. Datanode interfaces
The declaration of import API in datanode RPC:
service DataNode {
rpc Import(ImportTask) returns(common.Status) {}
}
6. Bulk Load task Assignment
There is a background knowledge that the inserted data shall be hashed into shards. Bulk load shall follow the same convention. There are two policy we can choose to satisfy this requirement:
- Assign the task to single Datanode. This datanode shall follow the same hashing rule and put the records into corresponding segments
- Assign the task to Datanode(s), which are responsible to watch the DmlChannels of the target Collection. Each datanode is responsible to handle its part of the data.
Considering the efficiency and flexibility, we shall implement option 1.
7. Result segments availability
By definition, the result segments shall be available altogether. Which means there shall be no intermediate state for loading.
To achieve this property, the segments shall be marked as "Importing" state and be invisible before the whole loading procedure completes.
8. Bulk Load with Delete
Constraint: The segments generated by Bulk Load shall not be affected by delete operations before the whole procedure is finished.
An extra attribute may be needed to mark the Load finish ts and all delta log before this ts shall be ignored.
9. Bulk Load and Query Cluster
After the load is done, the result segments needs to be loaded if the target collection/partition is loaded.
DataCoord has two ways to notify the Query Cluster there are some new segments to load
- Flush new segment
- Hand-off
Since we want to load the segments altogether, hand-off shall be a better candidate and some twist shall be taken:
- Allow adding segments without removing one
- Bring target segments online atomically.
10. Bulk Load and Index Building
The current behavior of query cluster is that if there is an index built for the collection, the segments will not be loaded(as sealed segment) before the index is built.
This constraint shall remain in first implementation of Bulk Load:
Constraint: The bulk load procedure shall include the period of index building of the result segments
11. Bulk Load as a tool
The bulk load logic can be extracted into a tool to run outside of Milvus process. It shall be implemented in the next release.
6 Comments
Xiaofan
{"table": {"rows": [{"id":1,"year":2021,"vector": [1.0,1.1,1.2]},{"id":2,"year":2022,"vector": [2.0,2.1,2.2]},{"id":3,"year":2023,"vector": [3.0,3.1,3.2]}]}}should we use collection and entity instead? also vector field should has a name? otherwise we are not able to support multi vector later ongroot
It is only an example, this format is not a strict rule.
Users can use different names to construct the JSON file like this:
{"my_collection": {"row_based_entities": [{"id":1,"year":2021,"vector1": [1.0,1.1,1.2],"vector2": [1.0,1.1,1.2]},{"id":2,"year":2022,"vector1": [2.0,2.1,2.2],"vector2": [1.0,1.1,1.2]},{"id":3,"year":2023,"vector1": [3.0,3.1,3.2],"vector2": [1.0,1.1,1.2]}]}}This json file contains two vector fields.
Then the "options" can be:
{"data_source": {......
},"external_data": {"target_collection":"TEST","chunks": [{"files": [{"path":"xxxx/file_1.json","type":"row_based","fields_mapping": {"my_collection.row_based_entities.id":"uid","my_collection.row_based_entities.year":"year","my_collection.row_based_entities.vector1":"vector1","my_collection.row_based_entities.vector2":"vector2"}}]}]}}Xiaofan
Numpy file is a binary format, we only treat it as vector data. Each Numpy file represents a vector field.
Maybe pandas dataframe is more similar to milvus data structure?
groot
Yes, each numpy file represents a vector field.
If there is a collection contains two vector fields, then the user needs to provide two numpy files and a json file for the "options":
"external_data": {"target_collection":"TEST","chunks": [{"files": [{"path":"xxxx/file_1.json","type":"column_based","fields_mapping": {"table.columns.id":"uid","table.columns.year":"year","table.columns.age":"age"}}]},{"files": [{"path":"xxxx/file_2.npy","fields_mapping": {"vector1":"vector1"}}]},{"files": [{"path":"xxxx/file_3.npy","fields_mapping": {"vector2":"vector2"}}]}]}Pandas dataframe is a csv-like format in memory, we can treat it as a kind of csv. We also can expand this api to support csv format:
"external_data": {"target_collection":"TEST","chunks": [{"files": [{"path":"xxxx/file_1.csv","fields_mapping": {"id":"uid","year":"year","age":"age"Xiaofan
gotca, so numpy only support vector data load right?
If you want to load scalar data then user has to use pandas frame or csv
groot
Yes, numpy only for vector data.
Vector data also can be store in json or csv. A customer has told us they use JSON to store scalar and vector data.
Scalar data can be json or csv.
At the first stage we support numpy and json.